Originally posted on Cracking the Enigma, January 2012
Having spent much of the past week struggling to make sense of my data, it’s good to come home, pour a glass of wine, put on some Sharon Jones, and, er… play with somebody else’s data!
Recently, I’ve discovered DataThief – an application that allows you to scan in a graph from a paper and extract the data points. Sometimes, this provides insights that really aren’t obvious from the original paper.
The other week, for example, I came across an intriguing neuroimaging study reported on the SFARI website. In the study, Judith Verhoeven and colleagues used diffusion tensor MRI to examine the superior longitudinal fasciculus, a bundle of nerve fibres that is assumed (although see this paper) to connect two brain regions involved in language production and comprehension – Broca’s area (left front-ish) with Wernicke’s area (left and back a bit).
Verhoeven et al. reported that integrity of the superior longitudinal fasciculus was compromised in kids with specific language impairment (SLI) – that is, kids who have language difficulties for no obvious reason. However, the same was not true of kids with autism, even though they had poorer language skills than those with SLI.
Taken at face value, this is a pretty major blow to the idea that autism and SLI have anything more than a superficial resemblance. DataThief, however, suggests otherwise.
The figure below is a scatterplot with each coloured shape representing a single child. On the x-axis is performance on a language test. On the y-axis is fractional anisotropy (FA) – the imaging measure used to assess the integrity of the left superior longitudinal fasciculus.
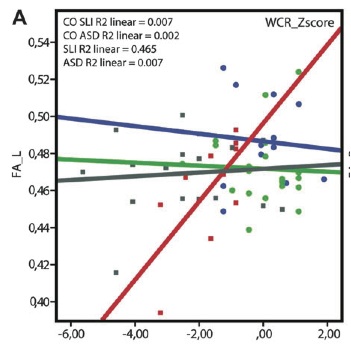
The purpose of the graph was to show the significant correlation between these two measures in the SLI group. But if we can read off the y-coordinates of each shape, we can show the distribution of fractional anisotropy scores for all three groups.
Cue DataThief.
It’s really just a case of clicking on three reference points for which you know the coordinates and then clicking on each of the data points in turn. Then you simply export the coordinates of the data points as a text file. The only thing I had to remember was to do the three groups separately so I knew which point belonged in which group.
Here’s the fractional anisotropy data replotted to show the distribution for each group. What we can now see is that there is a small subgroup of control kids who have really high FAs. There is also one autistic kid and one kid (arguably two) with SLI who have low FAs. Everyone else is pretty much in the middle.

On average, kids with SLI have lower than ‘normal’ fractional anisotropy [1], but looking at the spread of scores, you’d be hard pressed to conclude that this was a characteristic of SLI. Likewise, the overlap between the distributions of the autism and SLI groups is almost complete. Hardly evidence for fundamentally different neural mechanisms in the two disorders.
At the risk of sounding like a broken record, this once again highlights the importance of looking at individual variation within diagnostic groups such as autism and SLI, rather than (or as well as) looking at group averages.
But it also emphasizes a more general point (and this I have to stress is no criticism of the authors of this particular paper).
The data reported in a journal article are really just a snapshot of the actual data recorded, filtered through the authors’ preconceptions about what questions are interesting to ask and how to go about doing that. There’s an imperative to present the data in a neat, sanitized package, with all the rough edges and anomalies smoothed out; to tell a coherent story that will convince reviewers and editors that it’s worthy of publication in a reputable journal. Years of work and terabytes of data may be compressed into just two or three pages.
DataThief only takes us so far. It allows us to extract the information presented visually in the published article, but no further.
Most of the past week has been spent convincing myself that it doesn’t really matter how I analyse my data because the results come out the same regardless. This is reassuring for me, but it doesn’t mean that somebody else, looking at my data with fresh eyes and a different perspective, would not come to an entirely different set of conclusions.
In an ideal world, when a paper is published, researchers should also be able (and encouraged) to publish the data on which the paper is based, as well as the script showing exactly how those data were analysed.
There are, of course, many obstacles in the way and questions to be answered before this becomes standard practice. Who would host and maintain the data? Just how raw should the raw data be? What if the authors are writing multiple papers based on the same data set? Who gets credit for reanalyses of the data set? What happens if a reanalysis shows up an error in the original paper? If the research involves human participants, how do we reassure them that their anonymity will be maintained?
Undoubtedly, there are many more problems that I haven’t thought of. But, as scientists, we need to work through these issues and find ways to set our data free.
Reference:
Verhoeven, J., Rommel, N., Prodi, E., Leemans, A., Zink, I., Vandewalle, E., Noens, I., Wagemans, J., Steyaert, J., Boets, B., Van de Winckel, A., De Cock, P., Lagae, L., & Sunaert, S. (2011). Is There a Common Neuroanatomical Substrate of Language Deficit between Autism Spectrum Disorder and Specific Language Impairment? Cerebral Cortex DOI: 10.1093/cercor/bhr292
